Research
Uncovering complex evolutionary histories of somatic mutations
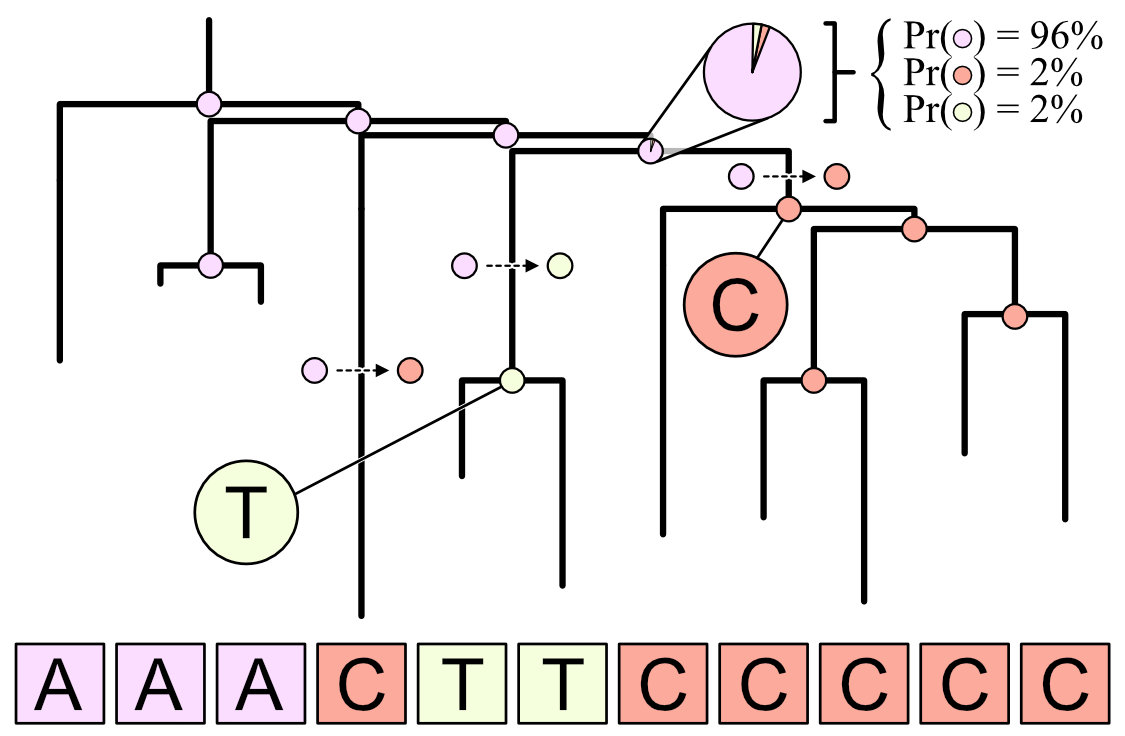
I lead the development and application of novel statistical methods aimed at studying somatic mutations that violate the infinite sites model of evolution, a simplifying assumption that asserts each mutation evolves only once within a population. By adopting a Bayesian approach for ancestral state reconstruction (called stochastic character mapping), my colleagues and I uncover previously hidden evolutionary complexity, including reversion and parallel mutation.
Example code:
Ongoing work: Somatic mutation SCM
Examining the role of telomere biology on hematopoiesis
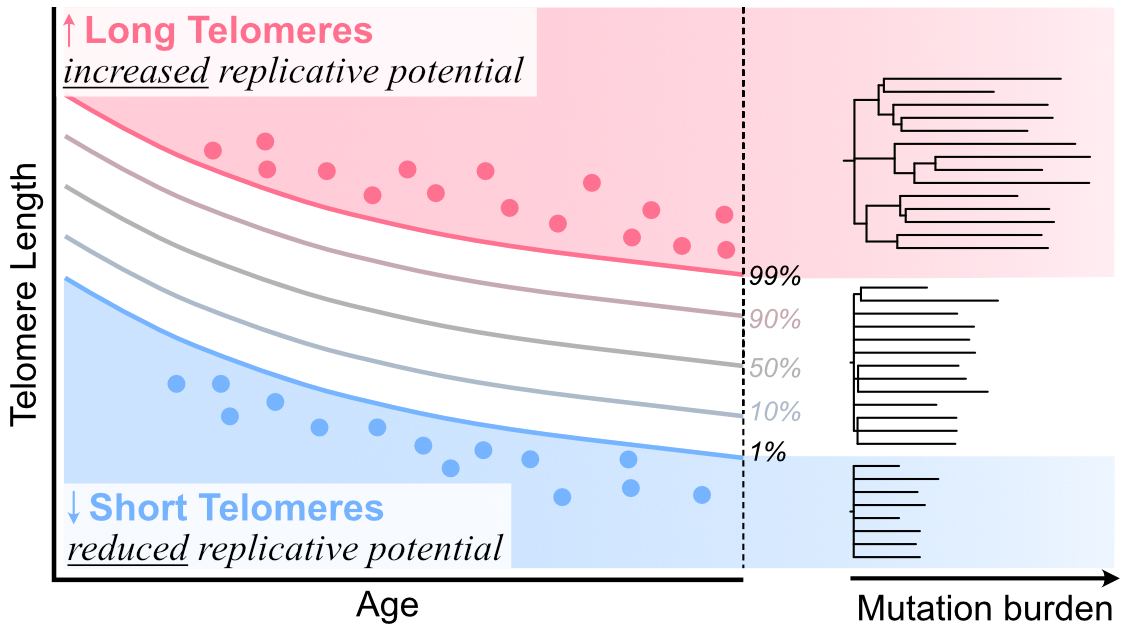
I am interested in understanding the evolutionary mechanisms through which heritable telomere length influences health and disease. My work in this field applies the principles of phylogenetics to examine the role of telomere shortening in governing hematopoiesis and leukemogenesis. We’ve shown that heritable long telomere length promotes cancer risk by increasing the proliferative lifetime of progenitors (DeBoy & Tassia et al. 2023, NEJM).
Example code:
Ongoing work: Short telomere length
DeBoy & Tassia et al. 2023: Long telomere length
Studying evolution across multiple scales of time

I am fascinated by the processes through which evolution manifests trait variation across multiple scales of time. In addition to my focal work studying the processes and outcomes of evolution within a single individual’s lifetime, I have also contributed to evolutionary studies at the scale of populations (e.g., Taylor et al. 2024, Nature) and species (e.g., Tassia et al. 2017, PNAS and Makova et al. 2024, Nature).
Example code:
Taylor et al. 2024: MAGE
Makova et al. 2024: Primate T2T assemblies of XY chromosomes